Uncertainty Quantification using Image Matching
Description
Based on paper and source code: - Sciacchitano, A., Wieneke, B., & Scarano, F. (2013). PIV uncertainty quantification by image matching. Measurement Science and Technology, 24 (4). https://doi.org/10.1088/0957-0233/24/4/045302. - http://piv.de/uncertainty/?page_id=221
Step 1: Particle peak detection
As described by (Sciacchitano et al., 2013, Eq. 1), the image intensity product \(\Pi\) from image matching intensities is defined as:
The peaks image \(\varphi\) is defined as:
Step 2: Disparity vector computation
The sub-pixel peak position estimator adopted here is the standard 3-point Gaussian fit. The particle positions of times \(t_1\) and \(t_2\) are defined as \(\boldsymbol{X}^1 = \{x^1_1,x^1_2,...,x^1_N\}\) and \(\boldsymbol{X}^2 = \{x^2_1,x^2_2,...,x^2_N\}\). Discrete disparity vectors are defined as:
Step 3: Disparity ensemble statistics inside window
The mean of disparity set inside a window is defined as:
where \(c_i = \sqrt{\Pi(x_i)}\) for \(i=1,2,...,N\).
The standard deviation of disparity set inside a window is defined as:
Finally, the instantaneous error (estimate) vector is defined as:
Setup
Packages
[1]:
%reload_ext autoreload
%autoreload 2
[2]:
import matplotlib.pyplot as plt
import numpy as np
import os
import pivuq
Load images
[3]:
parent_path = "./data/particledisparity_code_testdata/"
image_pair = np.array(
[plt.imread(os.path.join(parent_path + ipath)).astype("float") for ipath in ["B00010.tif", "B00011.tif"]]
)
Load reference velocity
[4]:
data = np.loadtxt(os.path.join(parent_path + "B00010_UQ.dat"), skiprows=3).T
I, J = 128, 128
X_ref = np.reshape(data[0], (I, J)) - 1 # zero-index
Y_ref = np.reshape(data[1], (I, J)) - 1
U_ref = np.stack((np.reshape(data[2], (I, J)), np.reshape(data[3], (I, J))))
e_ref = np.stack((np.reshape(data[4], (I, J)), np.reshape(data[5], (I, J))))
N_ref = np.reshape(data[6], (I, J))
Uncertainity quantificiation using image matching
[5]:
%%time
X, Y, delta, N, mu, sigma = pivuq.disparity.sws(
image_pair,
U_ref,
window_size=16,
grid_size=4,
window="gaussian",
radius=1,
sliding_window_subtraction=True,
ROI=[10, 450, 220, 430],
velocity_upsample_kind="linear",
warp_direction="center",
warp_order=-1,
warp_nsteps=1,
)
CPU times: user 535 ms, sys: 27.3 ms, total: 562 ms
Wall time: 274 ms
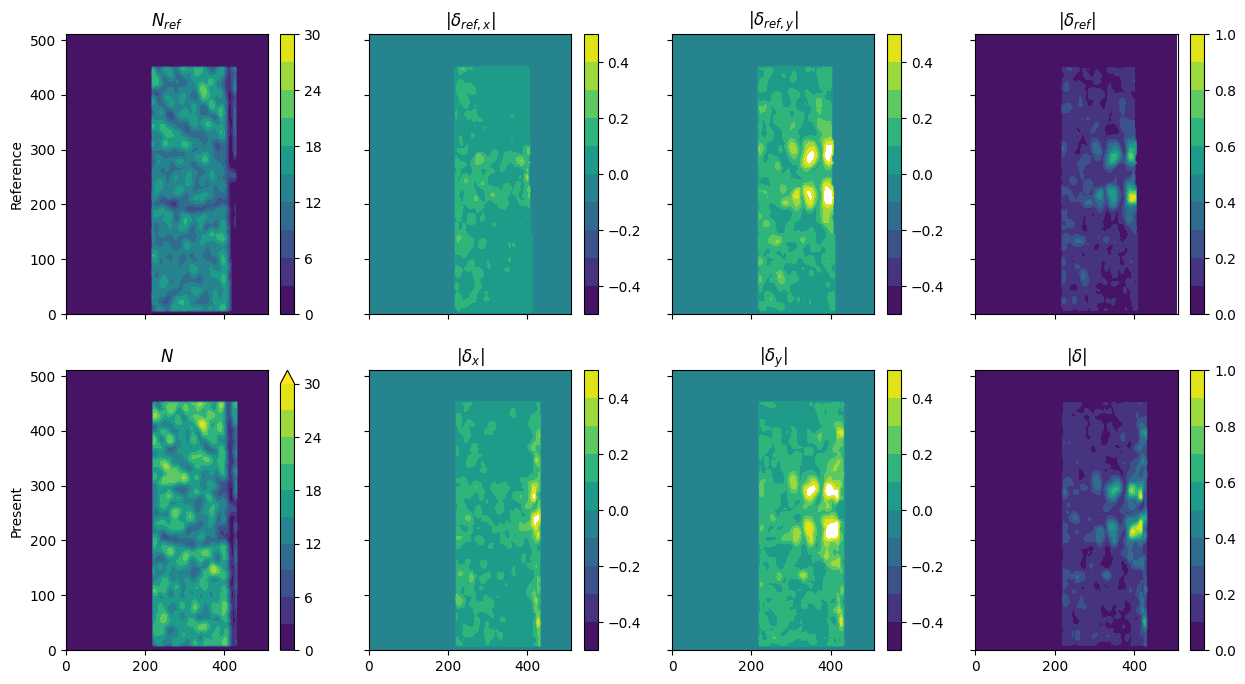
Plot: Instantaneous error map
[6]:
fig, axes = plt.subplots(nrows=2, ncols=4, sharex=True, sharey=True, figsize=(15, 8))
# References
ax = axes[0, 0]
im = ax.contourf(X_ref, Y_ref, N_ref, np.linspace(0, 30, 11))
fig.colorbar(im, ax=ax)
ax.set(title="$N_{ref}$", ylabel="Reference")
for i, (ax, var) in enumerate(zip(axes[0, 1:3], ["x", "y"])):
im = ax.contourf(X_ref, Y_ref, e_ref[i], np.linspace(-0.5, 0.5, 11))
fig.colorbar(im, ax=ax)
ax.set(title=f"$|\delta_{{ref,{var}}}|$")
ax = axes[0, 3]
im = ax.contourf(X_ref, Y_ref, np.linalg.norm(e_ref, axis=0), np.linspace(0, 1, 11))
fig.colorbar(im, ax=ax)
ax.set(title="$|\delta_{ref}|$")
# Present
ax = axes[1, 0]
im = ax.contourf(X, Y, N, np.linspace(0, 30, 11), extend="max")
fig.colorbar(im, ax=ax)
ax.set(title="$N$", ylabel="Present")
for i, (ax, var) in enumerate(zip(axes[1, 1:3], ["x", "y"])):
im = ax.contourf(X, Y, delta[i], np.linspace(-0.5, 0.5, 11))
fig.colorbar(im, ax=ax)
ax.set(title=f"$|\delta_{var}|$")
ax = axes[1, 3]
im = ax.contourf(X, Y, np.linalg.norm(delta, axis=0), np.linspace(0, 1, 11))
fig.colorbar(im, ax=ax)
ax.set(title="$|\delta|$");
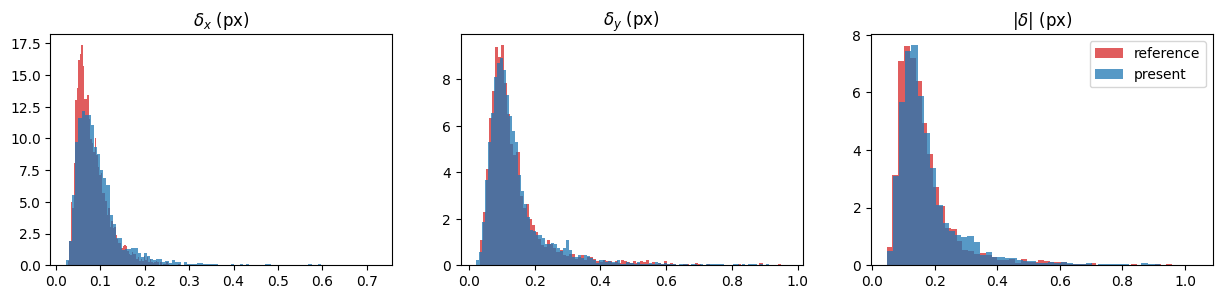
Plot: Instantaneous error histogram
[7]:
fig, axes = plt.subplots(ncols=3, figsize=(15, 3))
for i, (ax, label) in enumerate(zip(axes[:2], [r"$\delta_x$ (px)", r"$\delta_y$ (px)"])):
values = e_ref[i].ravel()
ax.hist(
values[np.abs(values) > 0],
bins=100,
density=True,
color="tab:red",
label="reference",
alpha=0.75,
)
ax.set(title=label)
for i, (ax, label) in enumerate(zip(axes[:2], [r"$\delta_x$ (px)", r"$\delta_y$ (px)"])):
values = delta[i].ravel()
ax.hist(
values[np.abs(values) > 0],
bins=100,
density=True,
color="tab:blue",
label="present",
alpha=0.75,
)
ax.set(title=label)
ax = axes[-1]
values = np.linalg.norm(e_ref, axis=0).ravel()
ax.hist(
values[np.abs(values) > 0],
bins=50,
density=True,
color="tab:red",
label="reference",
alpha=0.75,
)
values = np.linalg.norm(delta, axis=0).ravel()
ax.hist(
values[np.abs(values) > 0],
bins=50,
density=True,
color="tab:blue",
label="present",
alpha=0.75,
)
ax.set(title="$|\delta|$ (px)")
ax.legend();